Xavier Beltran Urbano

| Home |
|---|
| Projects |
| Publications |
| Awards and certifications |
| Services |
| Resume |
Projects
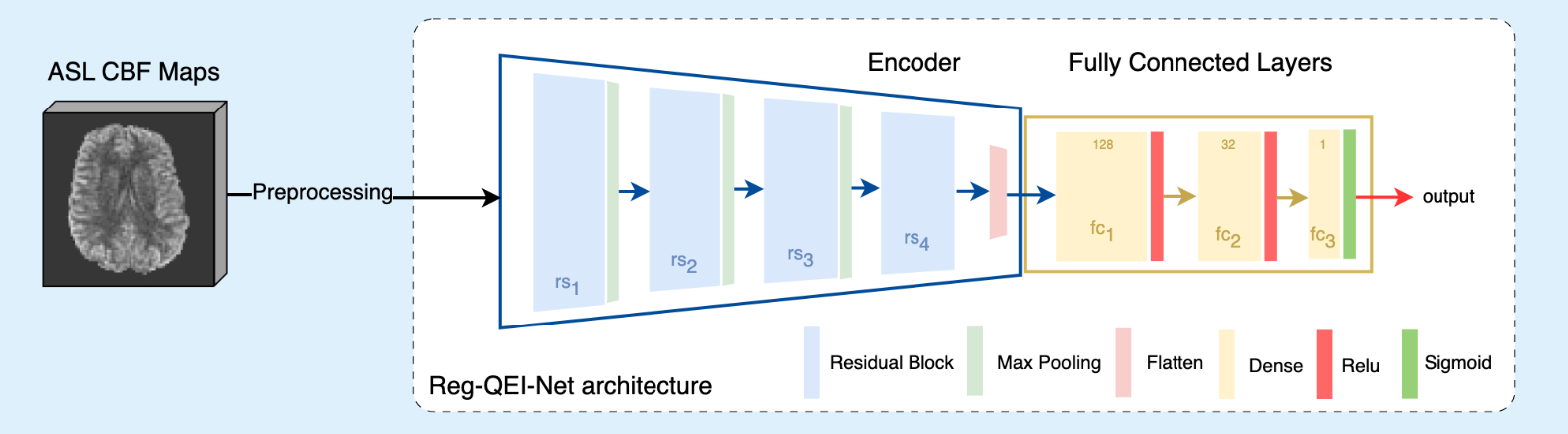
QEI-Net: A Deep learning-based automatic quality evaluation index for ASL CBF Maps
The project's goal is to develop and evaluate novel deep learning models to automate the quality control of ASL-derived cerebral blood flow (CBF) maps, providing continuous quality evaluation indices (QEIs). These models aim to enhance feature representation and performance, reducing the need for time-consuming, subjective manual assessments.
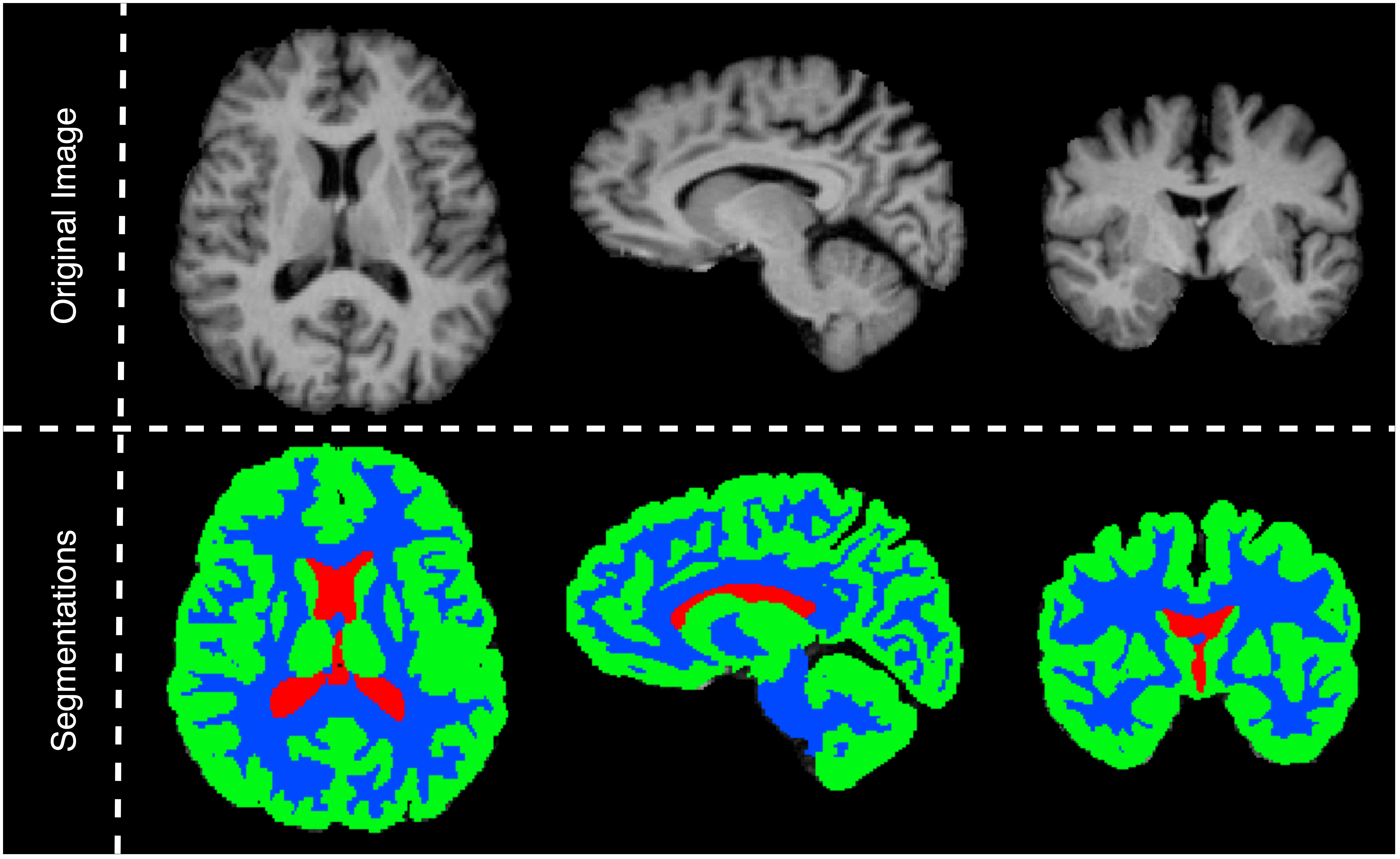
Enhancing Brain Tissue Analysis: A 2D and 3D CNN Ensemble Approach for MRI Segmentation
The project's main goal is to enhance brain tissue segmentation in MR imaging using an ensemble of 2D and 3D convolutional neural networks on the IBSR18 dataset. We achieved remarkable results, marked by Dice coefficients of 0.904 for cerebrospinal fluid (CSF), 0.945 for gray matter (GM), and 0.948 for white matter (WM).
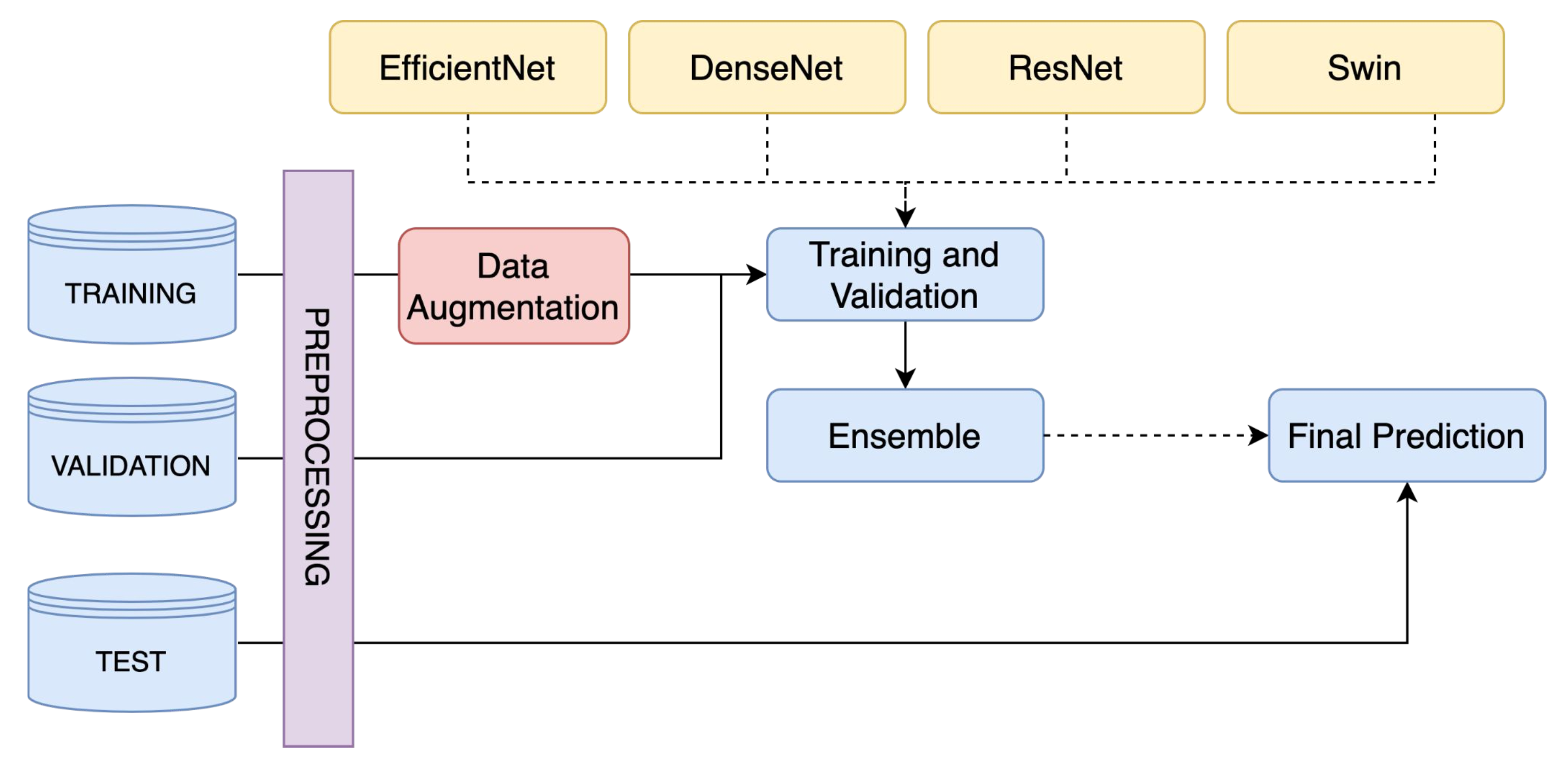
A Skin Lesion Classification Approach Using Deep Learning on the ISIC 2020
This project introduces a groundbreaking approach to skin lesion classification using deep learning, tackling two main challenges with impressive results. In binary classification, by ensembling EfficientNetB4, EfficientNetB5, and EfficientNetB6, an accuracy of 0.9336 was obtained. The multiclass classification challenge showed even greater success, where an ensemble model of EfficientB1, Swin S, Swin V2 S, and Swin V2 B achieved an accuracy of 0.9780 and a kappa value of 0.9604.
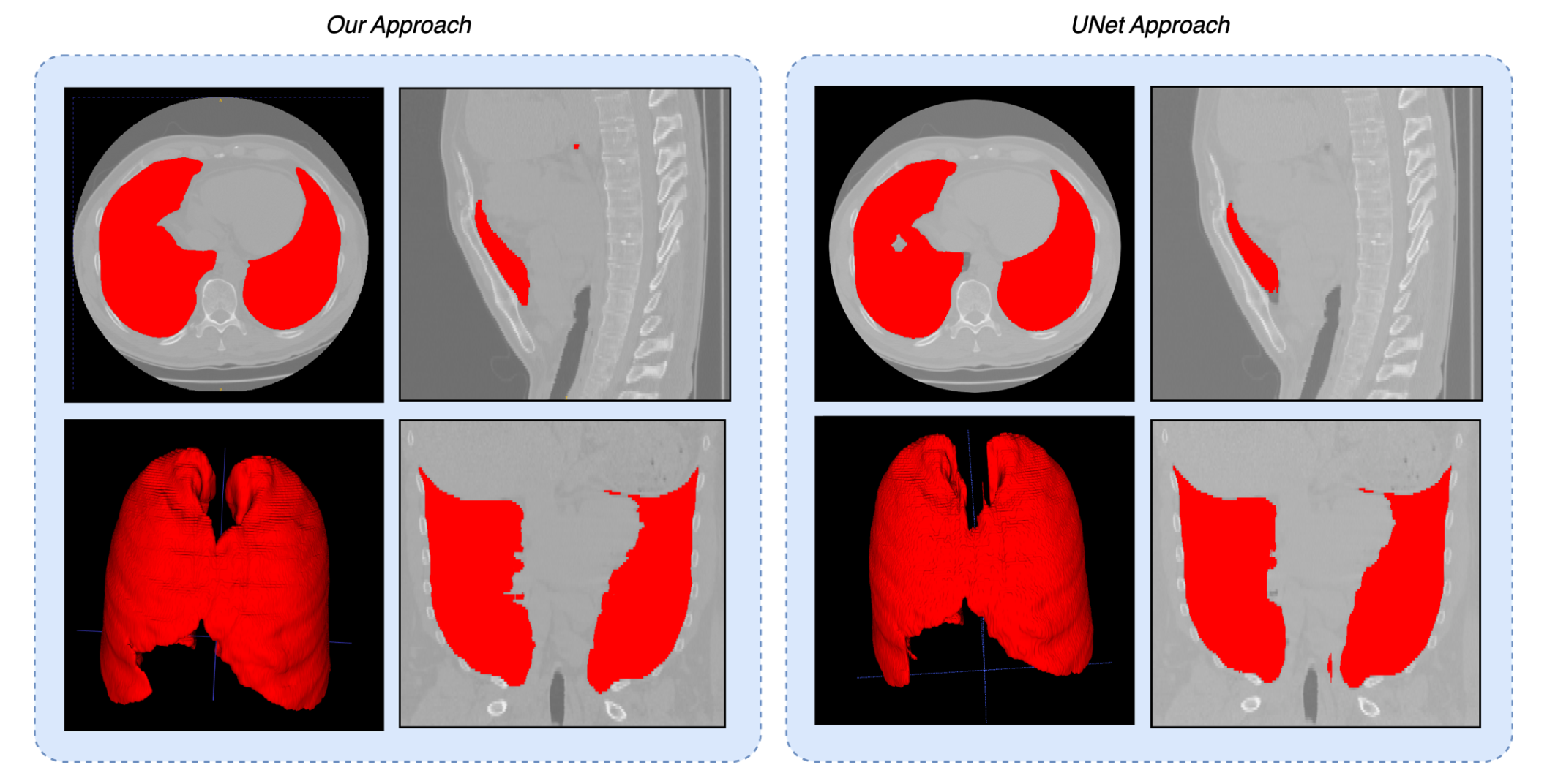
Image registration of chest CT volumes: 4DCT DIR-Lab Challenge
This project presents a comprehensive study on image registration of chest CT volumes, focusing on aligning thoracic structures across different respiratory phases. The results show itk-elastix, especially with custom parameter tuning and preprocessing, achieves a TRE of 2.63 ± 1.43 mm, significantly outperforming VoxelMorph, which yielded a TRE of 28.92±10.53 mm.
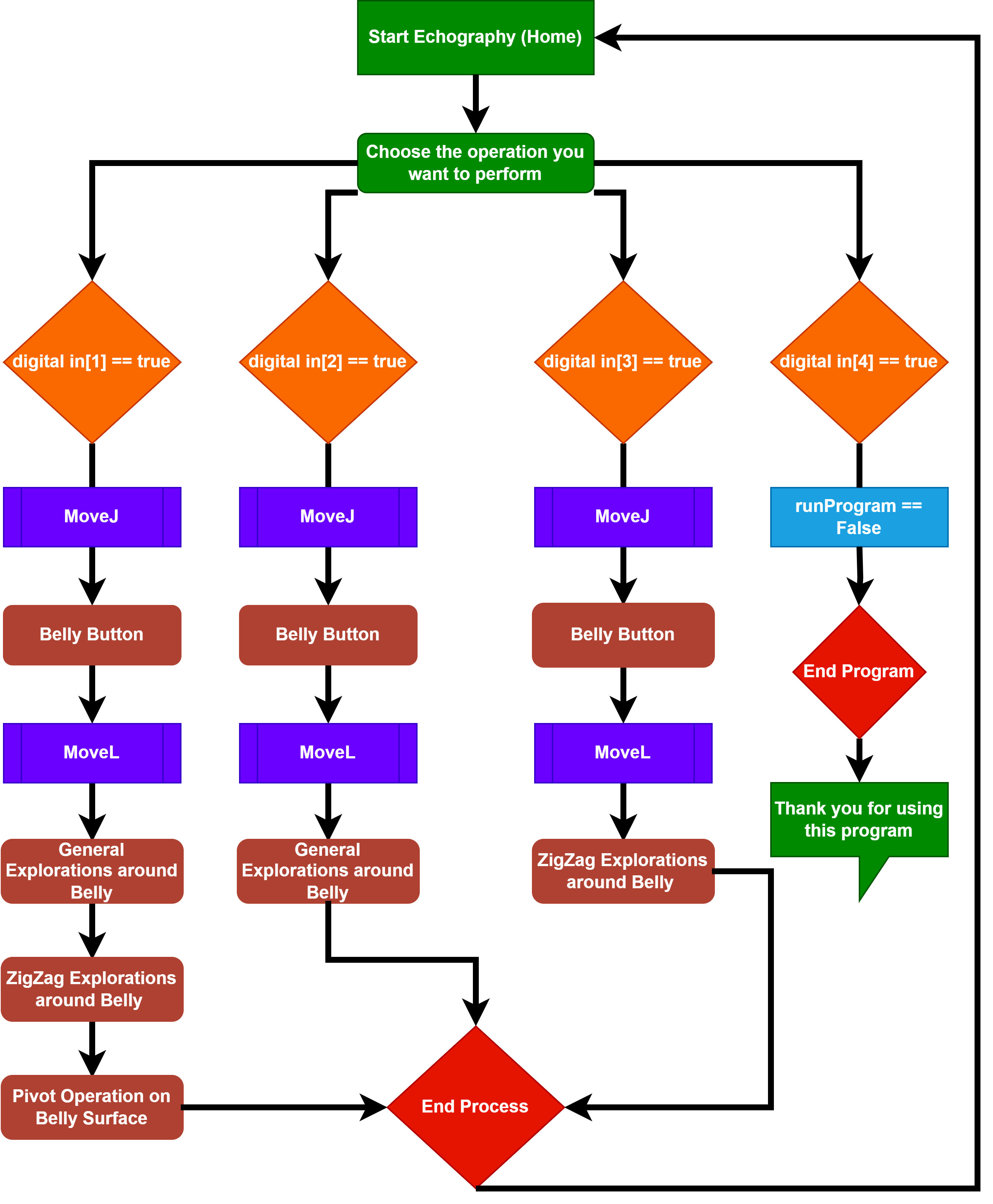
A Novel Automatic Ecography process for Pregnant Woman using a Collaborative Robot
This project introduces a novel automatic echography process for pregnant women, utilizing a collaborative robot equipped with an artificial transducer and a button control box for precise movement execution. It successfully automates standard echography movements - General Exploration, Zig-Zag, and Pivoting - ensuring safe practice with a controlled force application.
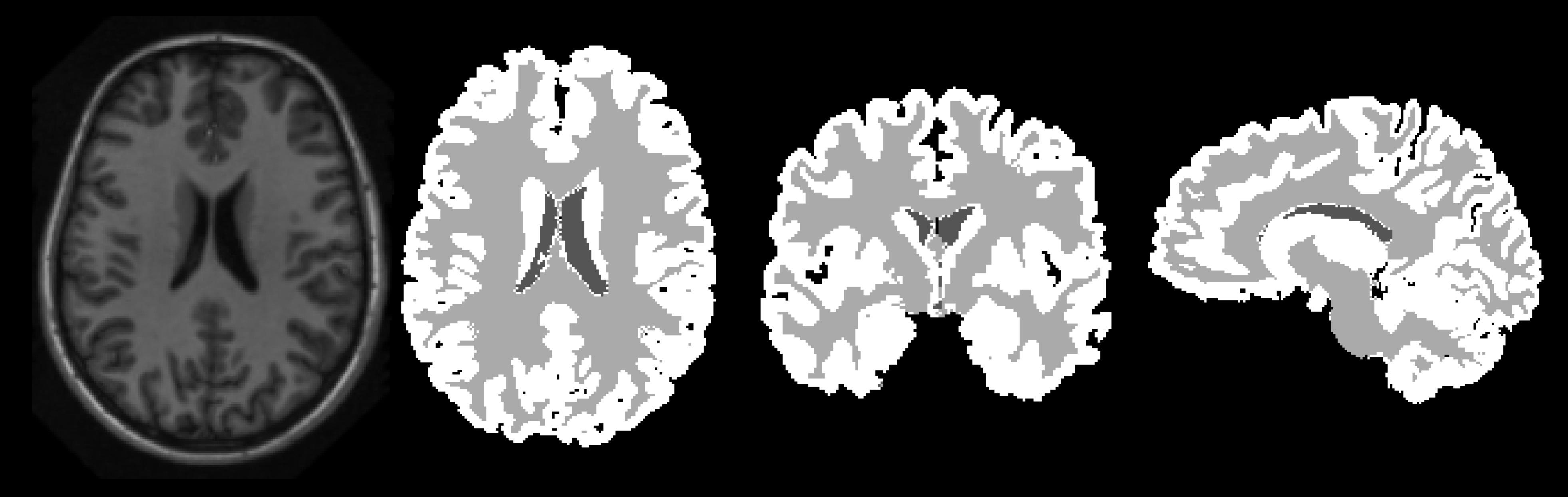
A Hybrid Approach for Brain Tissue Segmentation: Integrating Gaussian Mixture Models with Atlas-based and Tissue Modeling Techniques
The primary objective of this project is to enhance brain tissue segmentation by incorporating a probabilistic atlas with an intensity map into the GMM algorithm. The effectiveness of this integrated approach was rigorously evaluated, utilizing a combination of quantitative metrics and qualitative assessments to determine the performance of the segmentation strategies.
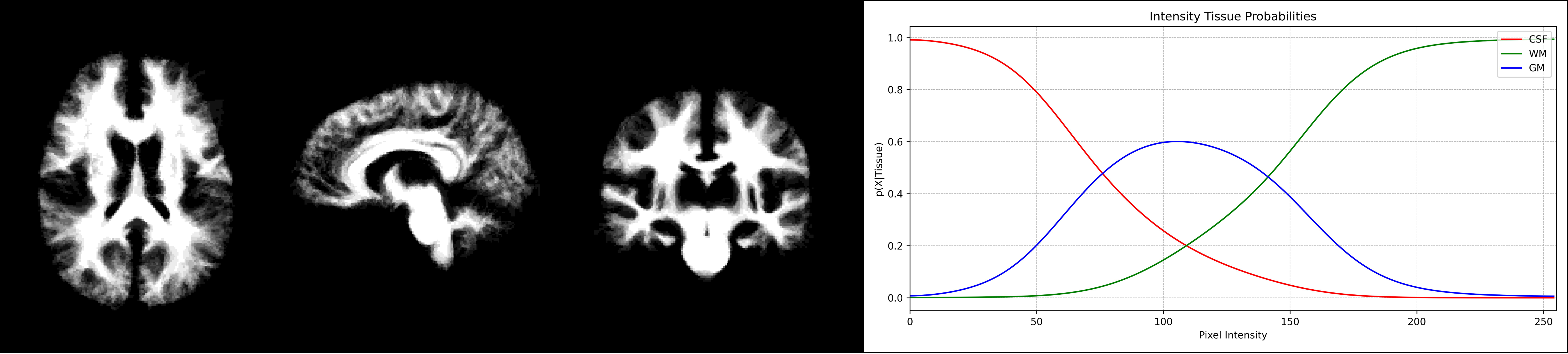
Development of a Probabilistic Brain Atlas and Tissue Probability Models
The objective of this project is to generate tissue probability models and a probabilistic brain atlas by utilizing MRI images. In order to establish a statistical distribution of brain tissues, it employs image registration techniques, including rigid, affine, and non-rigid transformations, to align images.
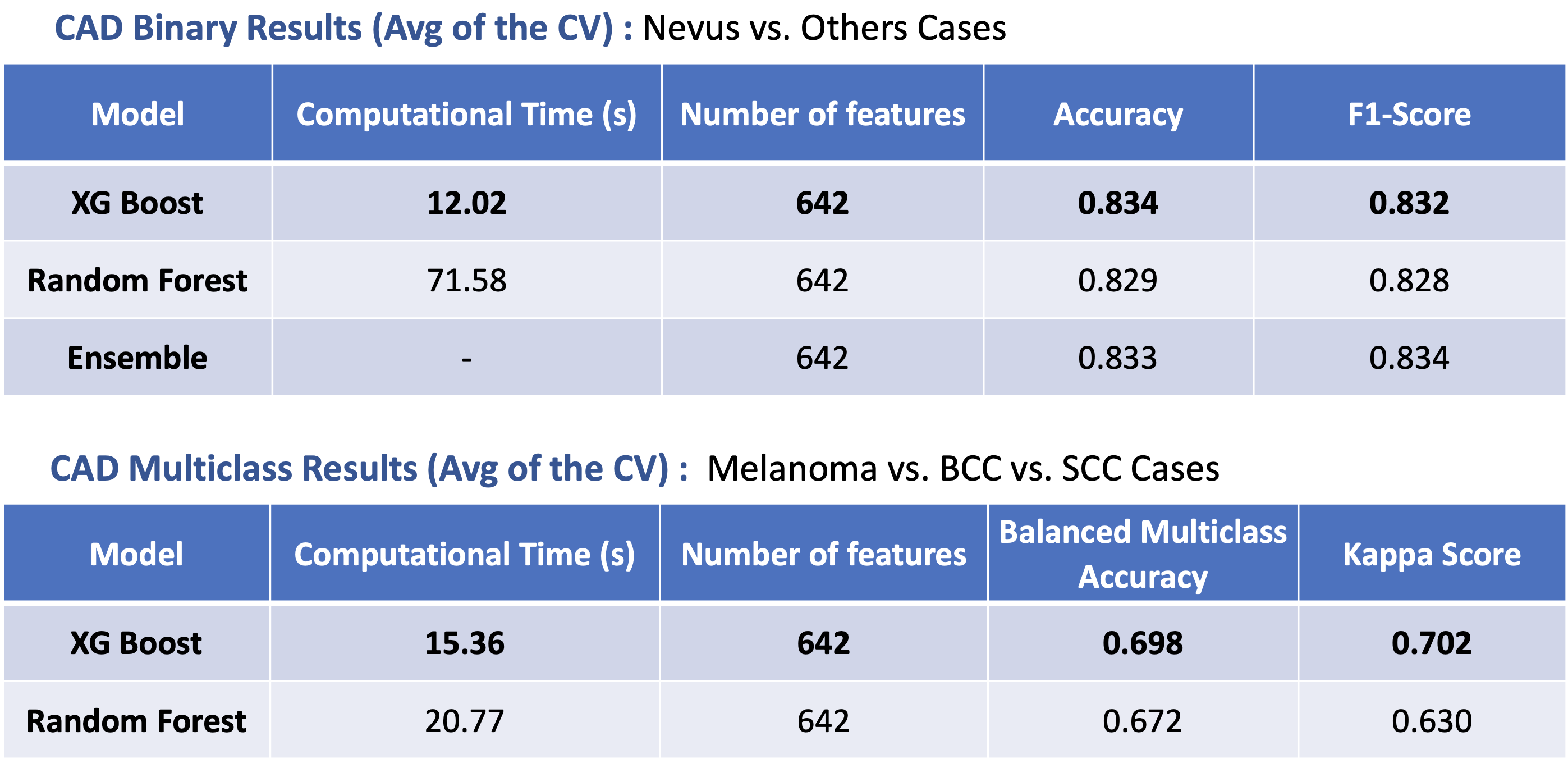
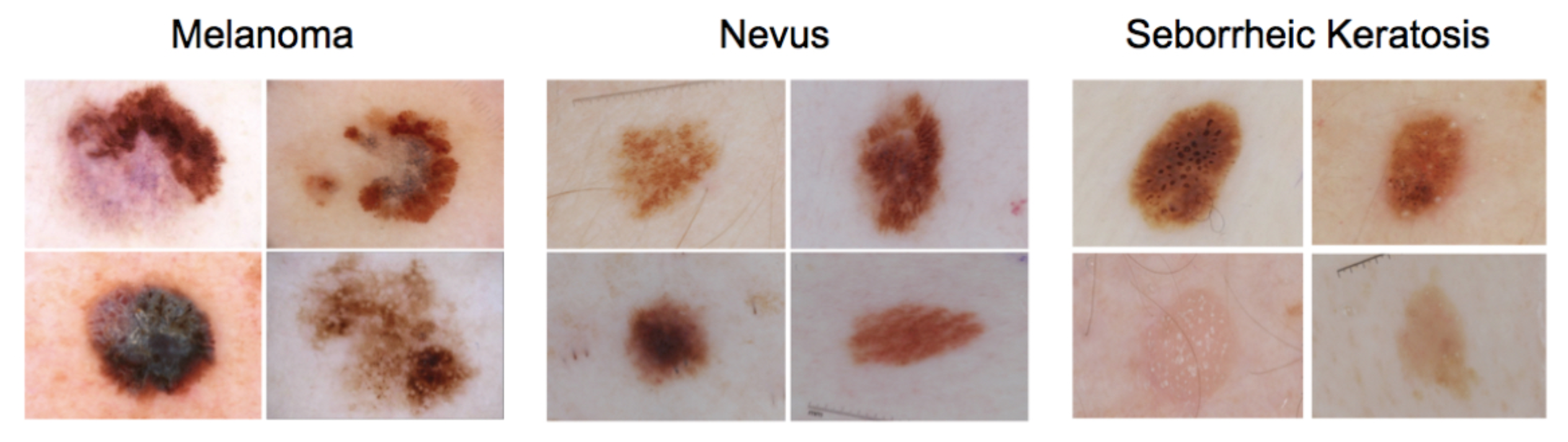
A Skin Lesion Classification Approach Using Traditional Machine Learning on the ISIC 2020 Dataset
The main goal of this project was to develop a Machine Learning approach to perform skin lesion classification using the ISIC 2020 Dataset. The results of this endevour demonstrated a robust performance, with an accuracy of 0.83 and a kappa score of 0.70 for binary and multiclass classification tasks, respectively.
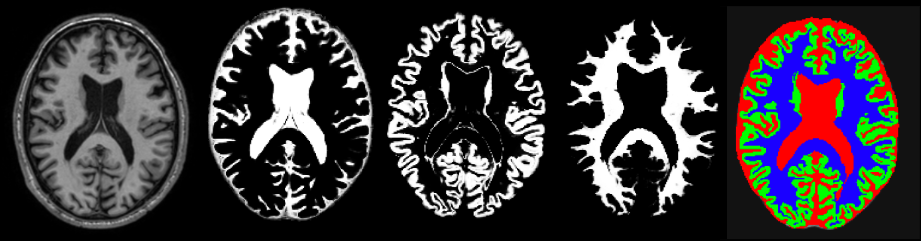
Brain tissue segmentation using Expectation Maximization (EM) algorithm for Gaussian Mixture Models (GMM)
In this report, the Expectation Maximization algorithm is combined with Gaussian Mixture Model to perform the task of brain tissue segmentation. The algorithm has been validated on a multi-modal dataset (n=5) reaching dice scores of 0.85, 0.80 and 0.81 for White Matter, Gray Matter and Cerebrospinal Fluid, respectively. Furthermore, the algorithm is compared to other segmentation tools such as K-Means or Statistical Parametric Mapping (SPM), achieving similar results.
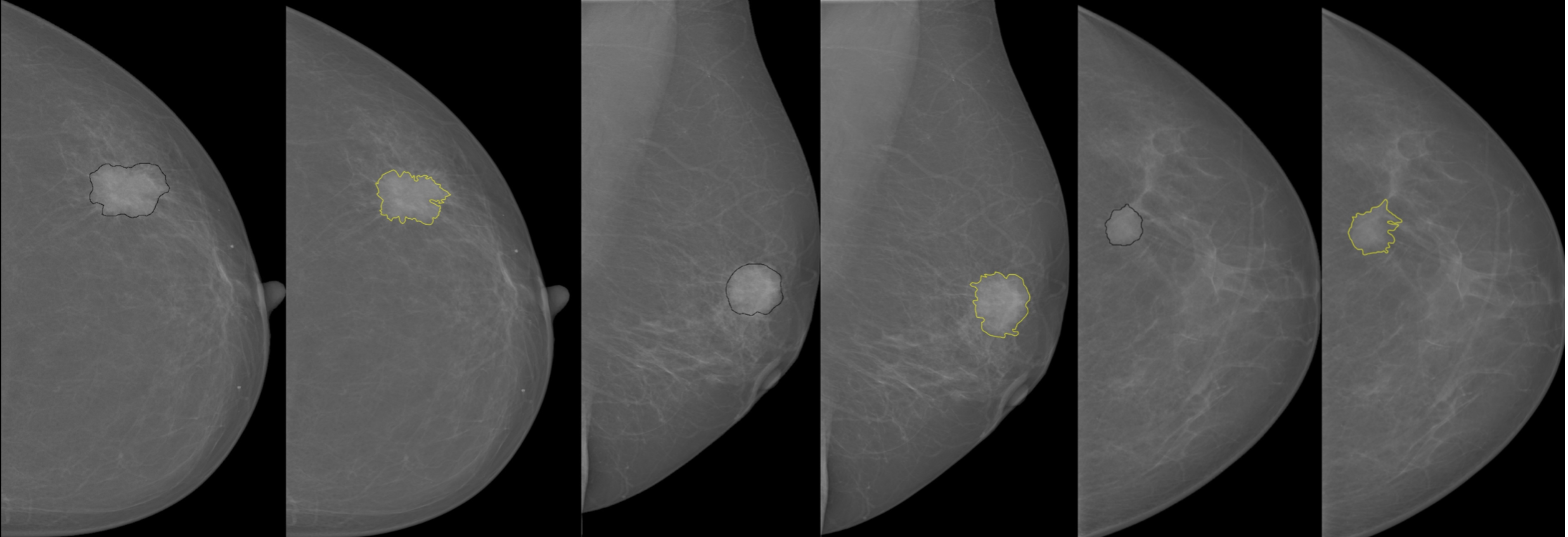
Breast Mass Detection System Based on a Multi-scale Approach
This project was implemented for mammographic mass detection and segmentation using a multi-scale morphological approach. This implementation was evaluated on InBreast dataset and was able to segment mass lesions with a sensitivity of 99.07%.
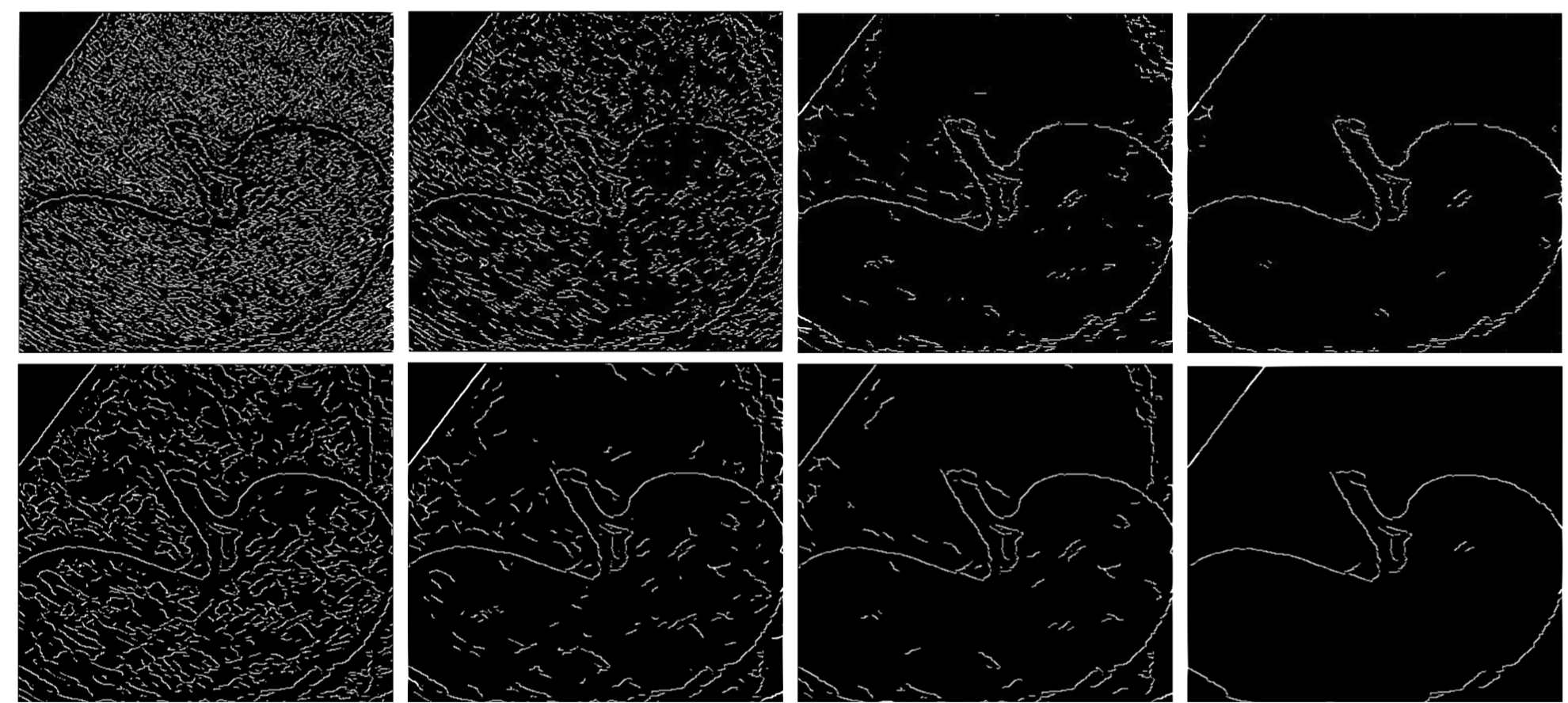

Edge Detection in Medical Ultrasound Images Using Adjusted Canny Edge Detection Algorithm
This project enhances edge detection in medical ultrasound images by modifying the traditional Canny Edge Detection Algorithm, substituting its Gaussian filter with a median filter and a dynamically weighted smoothing filter to more effectively reduce speckle noise, thereby improving the clarity and precision of internal organ boundaries.
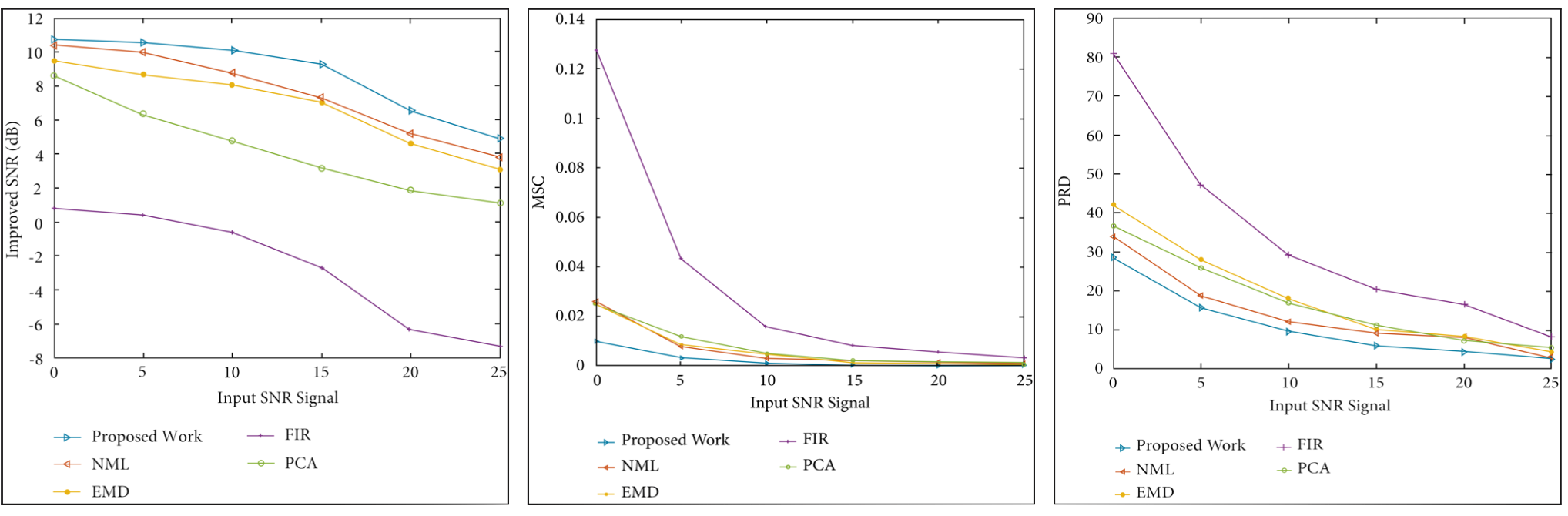
An Adaptive ECG Noise Removal Process Based on Empirical Mode Decomposition (EMD)
The research introduces an adaptive ECG noise removal technique using Empirical Mode Decomposition (EMD). By leveraging mean value adaptive biasing, the method improves signal quality and outperforms existing noise filtering approaches, offering enhanced accuracy in ECG signal analysis.